Note
New to DeepInverse? Get started with the basics with the 5 minute quickstart tutorial..
Scan-specific zero-shot SSDU for MRI#
We demonstrate scan-specific self-supervised learning, that is, learning to reconstruct MRI scans from a single accelerated sample without ground truth.
Here, we demonstrate fine-tuning a pretrained model (deepinv.models.RAM) [1]
with the weighted SSDU loss [2][3].
However, note that any of the self-supervised losses can be used to do this with varying performance [4].
For example see the example using Equivariant Imaging [5].
Note that, if more data is available, better results can be obtained by fine-tuning on more samples!
import torch
import deepinv as dinv
device = dinv.utils.get_device()
rng = torch.Generator(device=device).manual_seed(0)
rng_cpu = torch.Generator(device="cpu").manual_seed(0)
Selected GPU 0 with 4065.25 MiB free memory
Data#
First, download a demo single brain MRI volume (FLAIR sequence, SIEMENS Trio Tim 3T scanner) from the FastMRI brain dataset [6], via HuggingFace.
Important
By using this dataset, you confirm that you have agreed to and signed the FastMRI data use agreement.
DATA_DIR = dinv.utils.get_cache_home() / "datasets" / "fastMRI" / "multicoil_train"
SLICE_DIR = DATA_DIR / "slices"
DATA_DIR.mkdir(parents=True, exist_ok=True)
SLICE_DIR.mkdir(exist_ok=True)
dinv.utils.download_example("demo_fastmri_brain_multicoil.h5", DATA_DIR)
We use the FastMRI slice dataset provided in DeepInverse to load the volume and return all 16 slices.
The data is returned in the format x, y, params where params is a dictionary containing
the acceleration mask (simulated Gaussian mask with acceleration 6) and the
estimated coil sensitivity map.
Note
This loading takes a few seconds per slice, as it must estimate the coil sensitivity map on the fly.
dataset = dinv.datasets.FastMRISliceDataset(
DATA_DIR,
slice_index="all",
transform=dinv.datasets.MRISliceTransform(
mask_generator=dinv.physics.generator.GaussianMaskGenerator(
img_size=(256, 256), # this is overridden internally by true image size
acceleration=6,
center_fraction=0.08,
device="cpu",
rng=rng_cpu,
),
estimate_coil_maps=True,
),
)
0%| | 0/1 [00:00<?, ?it/s]
100%|██████████| 1/1 [00:00<00:00, 470.11it/s]
When training with data that is slow to be loaded, it is faster to save the pre-loaded slices:
Then the dataset can be loaded very quickly. We also pre-normalize to bring the data into a more friendly range. We also load a rough noise level as a param to be passed into the physics. The ground truth is loaded for evaluation later.
def loader(f):
x, y, params = torch.load(f, weights_only=True)
return x * 1e5, y * 1e5, params | {"sigma": 1e-5 * 1e5}
dataset = dinv.datasets.ImageFolder(SLICE_DIR, x_path="*.pt", loader=loader)
Physics#
The multicoil physics is defined every easily:
physics = dinv.physics.MultiCoilMRI(
device=device, noise_model=dinv.physics.GaussianNoise(0.0)
)
Model#
For the model we fine-tune a pretrained model (deepinv.models.RAM) [1]
Loss#
We define the weighted SSDU loss by first defining the generator that generates the splitting masks. These splitting masks are multiplied with the measurements as per the original paper. Finally, the weighted SSDU loss requires knowledge of the original physics generator to define the weight.
Tip
Feel free to use any self-supervised loss you like here!
split_generator = dinv.physics.generator.GaussianMaskGenerator(
(256, 256), acceleration=2, center_fraction=0.0, device=device
)
mask_generator = dinv.physics.generator.MultiplicativeSplittingMaskGenerator(
(256, 256), split_generator, device=device
)
physics_generator = dinv.physics.generator.GaussianMaskGenerator(
(256, 256), acceleration=6, center_fraction=0.04, rng=rng, device=device
)
loss = dinv.loss.mri.WeightedSplittingLoss(
mask_generator=mask_generator, physics_generator=physics_generator
)
Training#
We train the model using the self-supervised loss. We randomly split the volume into training slices and validation slices for early stopping (up to a maximum of 100 epochs). Because the FastMRI ground truth are cropped magnitude root-sum-of-squares reconstructions, we define a helper metric for evaluation later.
Hint
This example trains on GPU to accelerate training in the example.
def crop(x_net, x):
"""Crop to GT shape then take magnitude."""
return dinv.utils.MRIMixin().rss(
dinv.utils.MRIMixin().crop(x_net, shape=x.shape), multicoil=False
)
class CropPSNR(dinv.metric.PSNR):
def forward(self, x_net=None, x=None, *args, **kwargs):
return super().forward(crop(x_net, x), x, *args, **kwargs)
metric = CropPSNR(max_pixel=None)
train_dataset, val_dataset = torch.utils.data.random_split(
dataset, (0.8, 0.2), generator=rng_cpu
)
trainer = dinv.Trainer(
model=model,
physics=physics,
losses=loss,
metrics=metric,
optimizer=torch.optim.Adam(model.parameters(), lr=1e-6),
train_dataloader=torch.utils.data.DataLoader(
train_dataset, shuffle=True, generator=rng_cpu
),
eval_dataloader=torch.utils.data.DataLoader(val_dataset, generator=rng_cpu),
epochs=0 if str(device) == "cpu" else 100,
save_path=None,
early_stop=3,
early_stop_on_losses=True,
compute_train_metrics=False,
compute_eval_losses=True,
show_progress_bar=False, # disable progress bar for better vis in sphinx gallery.
device=device,
)
model = trainer.train()
model = model.eval()
/local/jtachell/deepinv/deepinv/deepinv/training/trainer.py:1356: UserWarning: non_blocking_transfers=True but DataLoader.pin_memory=False; set pin_memory=True to overlap host-device copies with compute.
self.setup_train()
The model has 35618813 trainable parameters
/local/jtachell/deepinv/deepinv/deepinv/physics/forward.py:978: UserWarning: At least one input physics is a DecomposablePhysics, but resulting physics will not be decomposable. `A_dagger` and `prox_l2` will fall back to approximate methods, which may impact performance.
warnings.warn(
/local/jtachell/deepinv/deepinv/deepinv/loss/mri/measplit.py:111: UserWarning: WeightedSplittingLoss detected new y width 213 in forward pass. Recalculating weight (this may take a second)...
warn(
Train epoch 0: TotalLoss=0.276
Eval epoch 0: TotalLoss=0.256, CropPSNR=30.676
Best model saved at epoch 1
Train epoch 1: TotalLoss=0.321
Eval epoch 1: TotalLoss=0.403, CropPSNR=30.72
Best model saved at epoch 2
Train epoch 2: TotalLoss=0.211
Eval epoch 2: TotalLoss=0.296, CropPSNR=30.765
Best model saved at epoch 3
Train epoch 3: TotalLoss=0.265
Eval epoch 3: TotalLoss=0.194, CropPSNR=30.814
Best model saved at epoch 4
Train epoch 4: TotalLoss=0.372
Eval epoch 4: TotalLoss=0.241, CropPSNR=30.844
Best model saved at epoch 5
Train epoch 5: TotalLoss=0.232
Eval epoch 5: TotalLoss=0.421, CropPSNR=30.866
Best model saved at epoch 6
Train epoch 6: TotalLoss=0.232
Eval epoch 6: TotalLoss=0.284, CropPSNR=30.889
Best model saved at epoch 7
Train epoch 7: TotalLoss=0.181
Eval epoch 7: TotalLoss=0.296, CropPSNR=30.914
Best model saved at epoch 8
Early stopping triggered at epoch 7 as validation metrics have not improved in the last 3 validation steps. Disable it with early_stop=None, or modify early_stop>0 to wait for more validation steps.
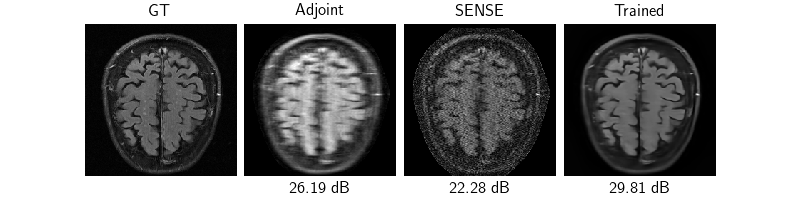
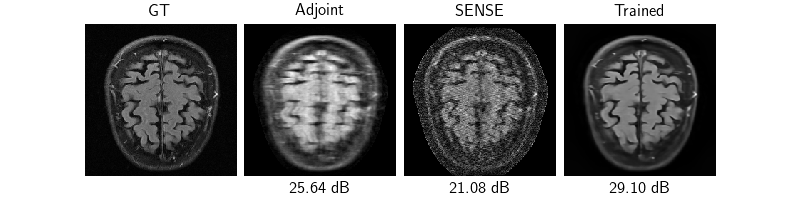
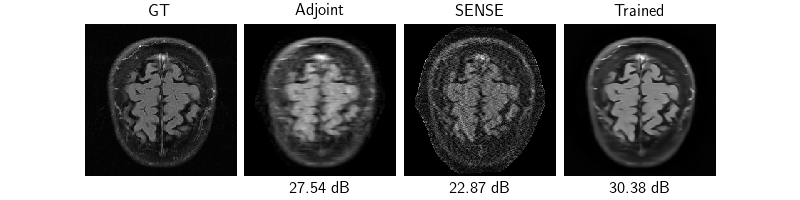
Evaluation#
Now that the model is trained, we test the model on 3 samples by evaluating the model, plotting and saving the reconstructions and evaluation metrics.
from torch.utils.data._utils.collate import default_collate
for i in [len(dataset) // 2 - 1, len(dataset) // 2, len(dataset) // 2 + 1]:
# Load slice
x, y, params = default_collate([dataset[i]])
x, y, params = (
x,
y.to(device),
{
k: (v.to(device) if isinstance(v, torch.Tensor) else v)
for (k, v) in params.items()
},
)
physics.update(**params)
# Compute baseline reconstructions
x_adj = physics.A_adjoint(y).detach().cpu()
x_dag = physics.A_dagger(y).detach().cpu()
# Evaluate model
with torch.no_grad():
x_hat = model(y, physics).detach().cpu()
dinv.utils.plot(
{
"GT": x,
"Adjoint": crop(x_adj, x),
"SENSE": crop(x_dag, x),
"Trained": crop(x_hat, x),
},
subtitles=[
"",
f"{metric(x_adj, x).item():.2f} dB",
f"{metric(x_dag, x).item():.2f} dB",
f"{metric(x_hat, x).item():.2f} dB",
],
)
- References:
Total running time of the script: (1 minutes 53.612 seconds)