Note
New to DeepInverse? Get started with the basics with the 5 minute quickstart tutorial..
Poisson Inverse Problems with Maximum-Likelihood Expectation-Maximization (MLEM)#
This example demonstrates how to solve Poisson inverse problems using the Maximum-Likelihood Expectation-Maximization (MLEM) algorithm Shepp and Vardi[1], also known as the Richardson-Lucy algorithm in the deconvolution setting Richardson[2], Lucy[3].
The Poisson observation model is:
where \(A\) is a linear forward operator, \(x \geq 0\) is the image to recover, \(\gamma > 0\) is the gain parameter, and \(\mathcal{P}\) denotes the Poisson distribution.
The MLEM algorithm solves the associated maximum-likelihood problem:
using the following iterative update rule:
where \(\odot\) denotes element-wise multiplication and the division is also element-wise. The MLEM algorithm is widely used in emission tomography such as Positron Emission Tomography (PET) and Single Photon Emission Computed Tomography (SPECT), where the Poisson noise model is a natural fit.
We show three scenarios of increasing complexity:
Deblurring with MLEM (no prior)
Deblurring with MLEM and Total-Variation (TV) prior
2D Computed Tomography (CT) with MLEM and TV prior
import torch
import deepinv as dinv
from pathlib import Path
from torchvision import transforms
from deepinv.utils.demo import load_dataset, load_example
from deepinv.utils.plotting import plot, plot_curves
Setup#
Set paths, device and random seed for reproducibility.
BASE_DIR = Path(".")
RESULTS_DIR = BASE_DIR / "results"
torch.manual_seed(0)
device = dinv.utils.get_freer_gpu() if torch.cuda.is_available() else "cpu"
Selected GPU 0 with 5015.25 MiB free memory
We use a single image from the Set3C dataset.
img_size = 128 if torch.cuda.is_available() else 64
val_transform = transforms.Compose(
[transforms.CenterCrop(img_size), transforms.ToTensor()]
)
dataset = load_dataset("set3c", transform=val_transform)
x = dataset[0].unsqueeze(0).to(device) # ground-truth image
Deblurring with MLEM without prior#
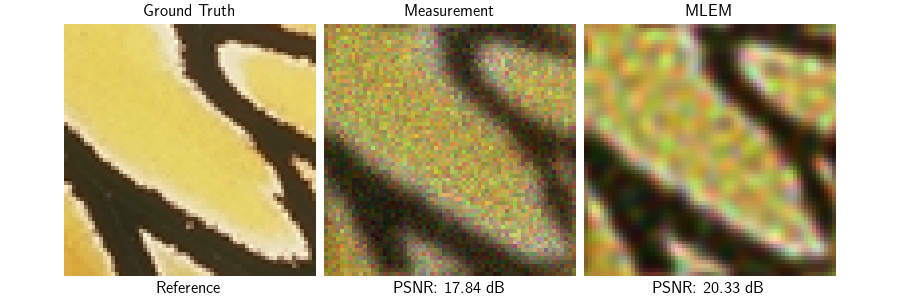
We create a Gaussian blur operator with Poisson noise. The MLEM/Richardson-Lucy algorithm is a standard approach for Poisson deconvolution without any prior.
# Define the blur kernel
n_channels = 3
filter_torch = dinv.physics.functional.gaussian_blur(sigma=(2, 2))
gain = 1 / 100
physics_blur = dinv.physics.BlurFFT(
img_size=(n_channels, img_size, img_size),
filter=filter_torch,
device=device,
noise_model=dinv.physics.PoissonNoise(
gain=gain, normalize=True, clip_positive=True
),
)
# Generate noisy blurred observation
y_blur = physics_blur(x)
The deepinv.optim.MLEM class wraps the MLEM iterations.
Without a prior, and in the case of deconvolution, this is equivalent to the classic Richardson-Lucy algorithm.
Note that without prior, the algorithm will create artifacts when noise is present in the observation.
data_fidelity = dinv.optim.PoissonLikelihood(gain=gain)
model_no_prior = dinv.optim.MLEM(
data_fidelity=data_fidelity,
prior=None,
max_iter=20,
early_stop=True,
thres_conv=1e-6,
crit_conv="residual",
verbose=True,
)
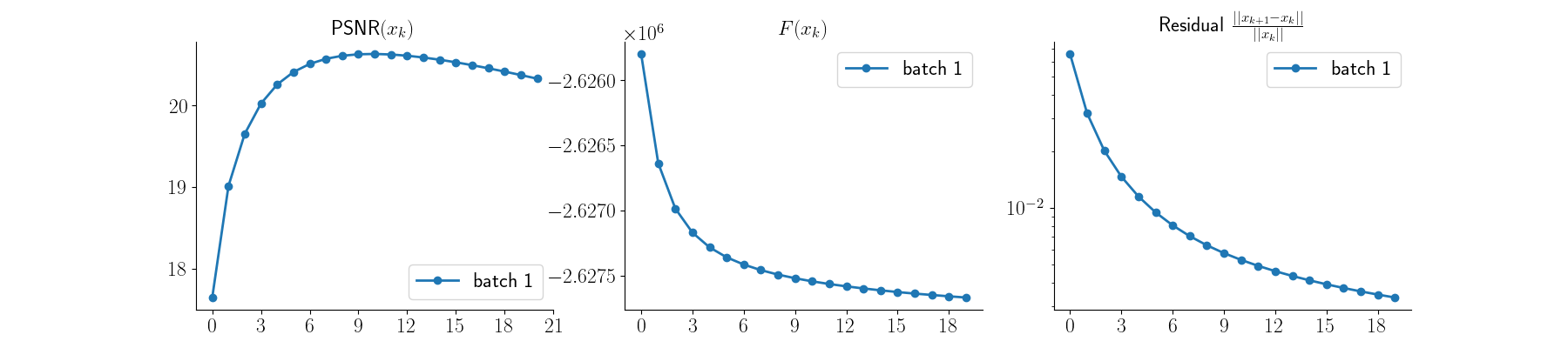
x_mlem, metrics_mlem = model_no_prior(
y_blur, physics_blur, x_gt=x, compute_metrics=True
)
Visualize results and PSNR values along with convergence curves
psnr_input = dinv.metric.PSNR()(x, y_blur)
psnr_mlem = dinv.metric.PSNR()(x, x_mlem)
plot(
{
"Ground Truth": x,
"Measurement": y_blur,
"MLEM": x_mlem,
},
subtitles=[
"Reference",
f"PSNR: {psnr_input.item():.2f} dB",
f"PSNR: {psnr_mlem.item():.2f} dB",
],
figsize=(9, 3),
)
plot_curves(metrics_mlem)
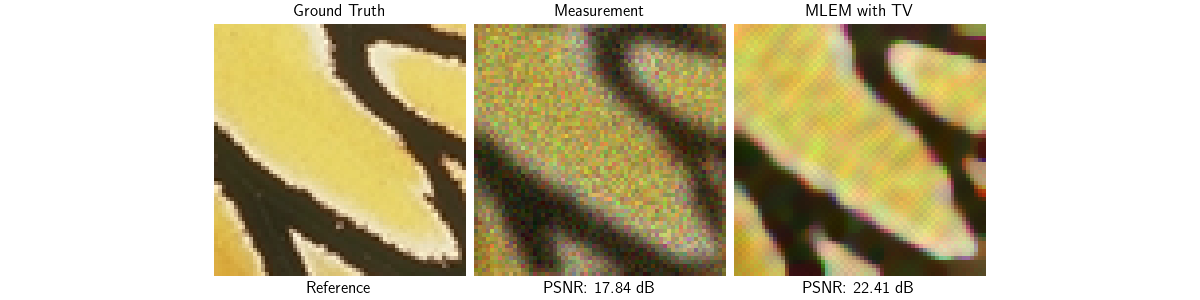
Deblurring with MLEM + TV prior#
As we saw, MLEM tends to amplify noise when no prior information is used. Adding a Total-Variation (TV) prior solves this issue while favoring piecewise constant solutions. There are several ways of modifying MLEM for regularized objectives: here we use the most straightforward approach called One-Step-Late (OSL) [4] which simply adds the gradient of the prior to the denominator of the MLEM update:
For non-smooth regularizations, the penalized MLEM update becomes:
where \(\partial \regname(x^k)\) is the subdifferential of the regularization at \(x^k\).
Any prior implementing the deepinv.optim.prior.Prior interface can be used in the deepinv.optim.MLEM class, and the proximal step is automatically computed when needed.
prior_tv = dinv.optim.prior.TVPrior(n_it_max=50)
model_tv = dinv.optim.MLEM(
data_fidelity=data_fidelity,
prior=prior_tv,
lambda_reg=0.02,
max_iter=100,
early_stop=True,
thres_conv=1e-6,
crit_conv="residual",
verbose=True,
)
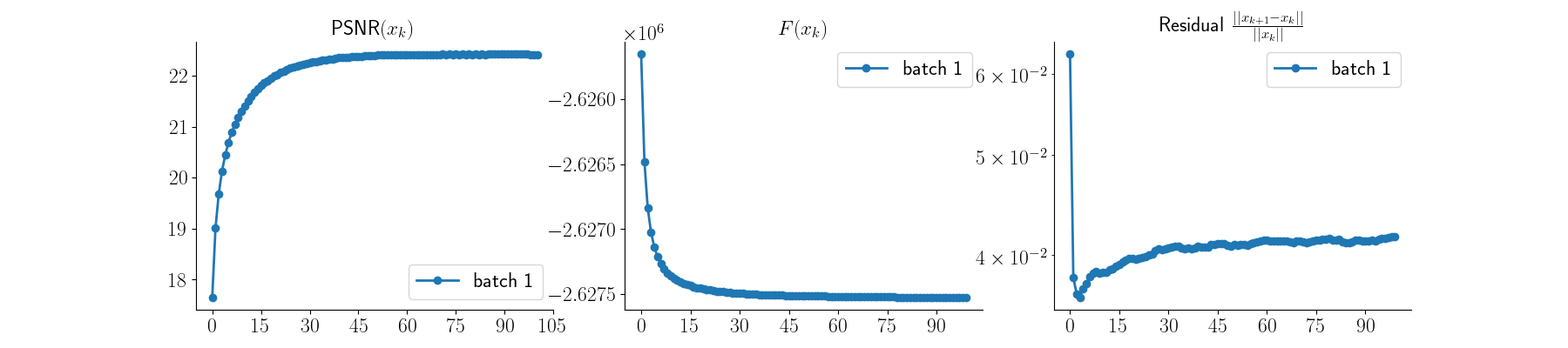
x_mlem_tv, metrics_mlem_tv = model_tv(
y_blur, physics_blur, x_gt=x, compute_metrics=True
)
Visualize results — MLEM + TV
psnr_mlem_tv = dinv.metric.PSNR()(x, x_mlem_tv)
plot(
{
"Ground Truth": x,
"Measurement": y_blur,
# "MLEM": x_mlem,
"MLEM with TV": x_mlem_tv,
},
subtitles=[
"Reference",
f"PSNR: {psnr_input.item():.2f} dB",
# f"PSNR: {psnr_mlem.item():.2f} dB",
f"PSNR: {psnr_mlem_tv.item():.2f} dB",
],
figsize=(12, 3),
)
plot_curves(metrics_mlem_tv)
Computed Tomography with MLEM + TV + custom metrics#
In emission tomography (PET/SPECT), the forward model is a Radon transform with Poisson statistics. Here we take the simple Shepp-Logan phantom as ground truth and use MLEM with TV prior to reconstruct it from its noisy sinogram.
# Load a grayscale image
val_transform_gray = transforms.Compose(
[
transforms.CenterCrop(img_size),
transforms.Grayscale(num_output_channels=1),
transforms.ToTensor(),
]
)
x_ct = load_example(
"SheppLogan.png",
img_size=img_size,
grayscale=True,
resize_mode="resize",
device=device,
)
Set up Tomography physics We define a parallel-beam tomography operator with 120 projection angles uniformly distributed between 0° and 180°, and Poisson noise.
num_angles = 120
gain_ct = 1 / 300
physics_ct = dinv.physics.Tomography(
img_width=img_size,
angles=num_angles,
device=device,
noise_model=dinv.physics.PoissonNoise(
gain=gain_ct, normalize=True, clip_positive=True
),
)
# Generate sinogram
y_ct = physics_ct(x_ct)
# Filtered back-projection as a simple baseline
x_fbp = physics_ct.A_dagger(y_ct)
/local/jtachell/deepinv/deepinv/deepinv/physics/tomography.py:187: UserWarning: The default value of `normalize` is not specified and will be automatically set to `True`. Set `normalize` explicitly to `True` or `False` to avoid this warning.
warn(
/local/jtachell/deepinv/deepinv/deepinv/physics/forward.py:487: UserWarning: Following torch.nn.Module's design, the 'device' attribute is deprecated and will be removed in a future version. To move the module's buffers/parameters to a different device, use the `to()` method.
warnings.warn(
Run MLEM + TV on the CT problem
data_fidelity_ct = dinv.optim.PoissonLikelihood(gain=gain_ct)
prior_tv_ct = dinv.optim.prior.TVPrior(n_it_max=50)
model_ct = dinv.optim.MLEM(
data_fidelity=data_fidelity_ct,
prior=prior_tv_ct,
lambda_reg=1e-2,
max_iter=50,
early_stop=True,
thres_conv=1e-6,
crit_conv="residual",
verbose=True,
)
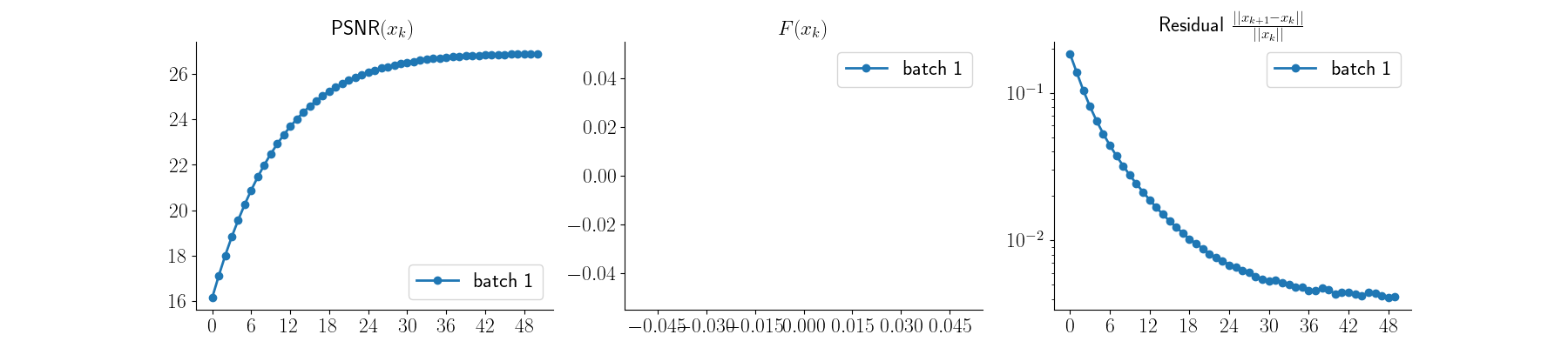
x_ct_recon, metrics_ct = model_ct(y_ct, physics_ct, x_gt=x_ct, compute_metrics=True)
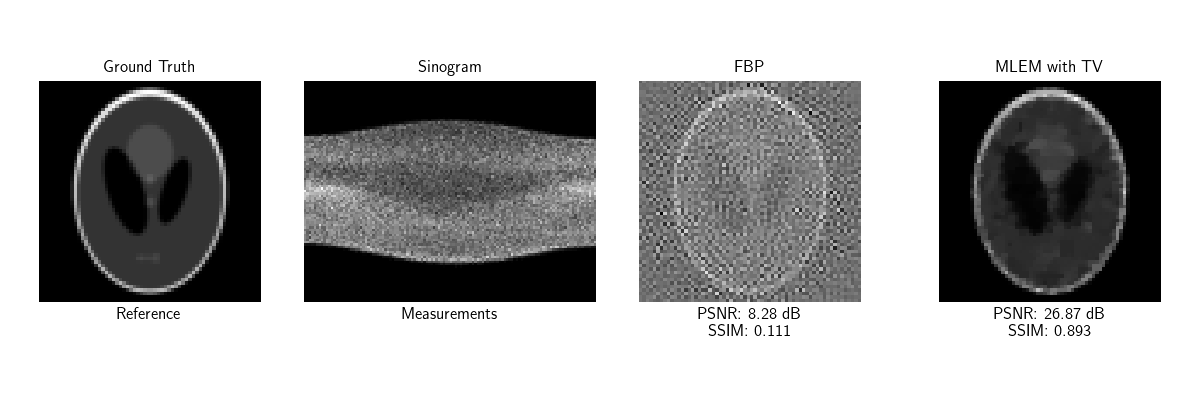
Visualize CT results and plot convergence curves
psnr_fbp = dinv.metric.PSNR()(x_ct, x_fbp)
psnr_ct = dinv.metric.PSNR()(x_ct, x_ct_recon)
ssim_fbp = dinv.metric.SSIM()(x_ct, x_fbp)
ssim_ct = dinv.metric.SSIM()(x_ct, x_ct_recon)
plot(
{
"Ground Truth": x_ct,
"Sinogram": y_ct,
"FBP": x_fbp,
"MLEM with TV": x_ct_recon,
},
subtitles=[
"Reference",
"Measurements",
f"PSNR: {psnr_fbp.item():.2f} dB\nSSIM: {ssim_fbp.item():.3f}",
f"PSNR: {psnr_ct.item():.2f} dB\nSSIM: {ssim_ct.item():.3f}",
],
figsize=(12, 4),
)
plot_curves(metrics_ct)
- References:
Total running time of the script: (0 minutes 5.988 seconds)