HDF5Dataset#
- class deepinv.datasets.HDF5Dataset(path, train=None, split=None, transform=None, load_physics_generator_params=False, dtype=torch.float, complex_dtype=torch.cfloat)[source]#
Bases:
ImageDatasetDeepInverse HDF5 dataset
DeepInverse features its own file format for imaging datasets designed as a subset of the HDF5 file format. A dataset in this format is typically obtained from a base dataset of ground-truth images and measured through a forward operator using the function
deepinv.datasets.generate_dataset(). This class features the code to load them in memory.
- Basics:
The file containing the dataset is opened in the constructor and remains opened until the method
close()is called.- Parameters:
path (str) – Path to the HDF5 file containing the dataset.
- Splits:
The dataset is structured in splits that are either freely named or understood specifically as training and testing splits. By convention, the training split is named
trainand the testing splittest. In both cases, the parametersplitcan be used to select one of the splits available in the dataset. For the specific case of training and testing splits, they can be loaded in using the boolean parametertrain. Iftrain=True, the training split is loaded, otherwise the testing split is loaded.By default, if neither
splitnortrainis provided, it attempts to load the training split. We don’t recommend relying on this behaviour which is likely to change in future versions of the library.Warning
If both
splitandtrainare provided, thensplittakes precedence andtrainis ignored. We recommend that you only specify one of the two parameters to avoid errors.Note
A single instance of the class holds a single split of the dataset. If you wish to load multiple splits, you must instantiate the class once per split. See for instance Training a reconstruction model.
- Parameters:
split (str, None) – The name of the split to load, for instance
"train","test"or"val"`. It can be left unspecified iftrainis used instead.train (bool) – If
splitis left unspecified, uses"train"as the split name if set toTrueand"test"otherwise. Note that ifsplitis specified, this parameter is ignored with a warning.
- Entries:
HDF5 datasets adhere to our conventions for datasets. In particular, their entries are either pairs of ground truth images and measuements
(x, y)or triplets with additional physics parameters(x, y, params). It is possible that the datast does not contain ground truth data and in this case the ground truth is replaced by a scalar NaN tensor.Physics parameters represent additional information about the measurement process. For instance the mask for inpainting or the blur kernel for deblurring. HDF5 datasets can contain the physics parameters used to generate each set of measurements and in that case, they are returned with each entry as a dictionary as long as the parameter
load_physics_generator_paramsis set toTrue. Note that if the parameter is set and the dataset does not contain any physics parameter, an empty dictionary is returned nonetheless.Measurements intended to be used with
deepinv.physics.StackedPhysicsare stored across multiple members of the HDF5 file, one per operator. In that case, member names follow the formaty{i}_{split_name}whereidenotes the stack index (starting at 0) and they are loaded as adeepinv.utils.tensorlist.TensorList.Note
Physics parameters are identified using a fallback logic. Namely, every member with a name of the form
{prefix}_{split_name}that is neither interpreted as containing ground truths or measurements (including stacked measurements) defaults to being interpreted as containing physics parameter, withprefixdenoting the parameter name. In particular, the joint presence of physics parameters and stacked measurements is generally supported as long as custom physics parameter names cannot be misinterpreted as ground truth or measurements names, for instancex,y, andy0are unsupported parameter names.Note
HDF5 datasets always contain measurements even though our conventions permit datasets with only ground truths (with or without physics parameters).
- Parameters:
load_physics_generator_params (bool) – Return the physics parameters with each entry. If no physics parameter is featured in the dataset, an empty dictionary is returned nonetheless.
- Pre-processing:
The data loaded in from the disk is not necessarily returned as is. The pre-processing pipeline contains two steps. First, the real and complex numbers are cast to user-provided dtypes using the parameters
dtypeandcomplex_dtype. Then, an optional transform provided by the user through the parametertransformis applied to the ground truth image.Note
The user-provided transformation is only applied to the ground truth image. It does not affect the measurements or the physics parameters.
- Parameters:
dtype (torch.dtype, str) – The dtype for real-valued numbers, by default
torch.float.complex_dtype (torch.dtype, str) – The dtype for complex-valued numbers, by default
torch.cfloat.transform (Transform, Callable, None) – An optional transformation applied to the ground truth.
Examples using HDF5Dataset:#

Imaging inverse problems with adversarial networks

Regularization by Denoising (RED) for Super-Resolution.
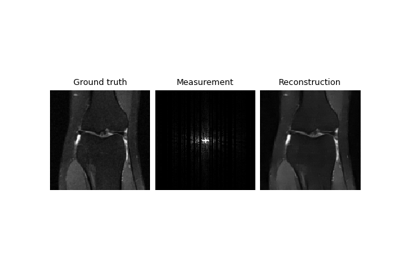
Self-supervised learning with Equivariant Imaging for MRI.

Self-supervised learning with Equivariant Splitting

Self-supervised learning from incomplete measurements of multiple operators.

Self-supervised denoising with the Neighbor2Neighbor loss.

Self-supervised denoising with the Generalized R2R loss.

Self-supervised learning with measurement splitting

Deep Equilibrium (DEQ) algorithms for image deblurring

Learned Iterative Soft-Thresholding Algorithm (LISTA) for compressed sensing

Unfolded Chambolle-Pock for constrained image inpainting